Setup code
We benchmark RmlxStats against base R and specialized fast fitting
packages, across varying numbers of cases (n) and
predictors (p). We also check accuracy.
Benchmarking was run on an M2 Macbook Air.
| Metadata | Metadata |
|---|---|
| Generated on | 2026-05-22 |
| Commit | 03bcaaf |
| Branch | master |
| Rmlx version | 0.4.0 |
Benchmarking Code
Data Generation
Code
set.seed(20251111)
n_sizes <- c(10000, 50000, 250000)
p_sizes <- c(50, 100, 200, 400, 800)
n_max <- max(n_sizes)
p_max <- max(p_sizes)
X <- matrix(rnorm(n_max * p_max), nrow = n_max, ncol = p_max)
colnames(X) <- paste0("x", seq_len(p_max))
X_glmnet <- matrix(rnorm(n_max * p_max), nrow = n_max, ncol = p_max)
for (j in 2:p_max) {
X_glmnet[, j] <- 0.6 * X_glmnet[, j - 1] + sqrt(1 - 0.6^2) * X_glmnet[, j]
}
colnames(X_glmnet) <- paste0("x", seq_len(p_max))
beta_true <- rnorm(p_max, mean = 0, sd = 0.5)
y_continuous <- drop(X %*% beta_true + rnorm(n_max, sd = 5))
# only 1 in 10 predictors matter:
beta_glmnet_true <- rep(0, p_max)
beta_glmnet_true[seq(10, p_max, 10)] <- rnorm(length(seq(10, p_max, 10)), sd = 0.35)
y_sparse <- drop(X_glmnet %*% beta_glmnet_true + rnorm(n_max, sd = 5))
linpred <- drop(X %*% beta_true) / 10
prob <- 1 / (1 + exp(-linpred))
y_binary <- rbinom(n_max, size = 1, prob = prob)
full_data <- data.frame(
y_cont = y_continuous,
y_bin = y_binary,
y_sparse = y_sparse,
X
)
# for fast debugging
if (params$develop) {
n_sizes <- n_sizes/10
p_sizes <- p_sizes/5
}
bench_grid <- expand.grid(
n = n_sizes,
p = p_sizes,
stringsAsFactors = FALSE
)
bench_grid <- bench_grid[bench_grid$n > bench_grid$p, ]
all_results <- data.frame()
all_accuracy <- data.frame()Helper functions
relative_rmse <- function(estimate, truth) {
scale <- sqrt(mean(truth^2))
sqrt(mean((estimate - truth)^2)) / max(scale, 1e-12)
}
safe_relative_rmse <- function(estimate, truth) {
estimate <- as.numeric(estimate)
truth <- as.numeric(truth)
if (anyNA(estimate) || anyNA(truth) ||
any(!is.finite(estimate)) || any(!is.finite(truth))) {
return(NA_real_)
}
relative_rmse(estimate, truth)
}
make_pca_fixture <- function(n, p, rank_k, noise_sd = 2) {
scores_raw <- qr.Q(qr(matrix(rnorm(n * rank_k), nrow = n, ncol = rank_k)))
scores_raw <- scale(scores_raw, center = TRUE, scale = FALSE)
scores_basis <- qr.Q(qr(scores_raw))
rotation <- qr.Q(qr(matrix(rnorm(p * rank_k), nrow = p, ncol = rank_k)))
sdev_true <- seq(from = 2.5, to = 1.5, length.out = rank_k)
singular_values <- sdev_true * sqrt(n - 1)
x_signal <- scores_basis %*% diag(singular_values, nrow = rank_k) %*%
t(rotation)
x <- x_signal + matrix(rnorm(n * p, sd = noise_sd), nrow = n, ncol = p)
colnames(x) <- paste0("pcx", seq_len(p))
list(
x = x,
rotation = rotation,
sdev = sdev_true,
rank = rank_k
)
}
extract_pca_rotation <- function(fit) {
if (inherits(fit, "big_SVD")) {
fit$v
} else {
fit$rotation
}
}
extract_pca_sdev <- function(fit) {
if (inherits(fit, "big_SVD")) {
fit$d / sqrt(nrow(fit$u) - 1)
} else {
fit$sdev
}
}
projector_error <- function(estimate, truth) {
estimate <- as.matrix(estimate)
truth <- as.matrix(truth)
proj_estimate <- estimate %*% t(estimate)
proj_truth <- truth %*% t(truth)
sqrt(sum((proj_estimate - proj_truth)^2)) / sqrt(sum(proj_truth^2))
}
pca_accuracy_score <- function(fit, truth_rotation, truth_sdev) {
projector_error(extract_pca_rotation(fit), truth_rotation) +
relative_rmse(
as.numeric(extract_pca_sdev(fit)[seq_along(truth_sdev)]),
truth_sdev
)
}
make_bootstrap_data <- function(n, p) {
x <- X[1:n, 1:p, drop = FALSE]
colnames(x) <- paste0("x", seq_len(p))
beta <- beta_true[seq_len(p)]
sigma <- 1 + 2 * abs(x[, 1])
y <- drop(x %*% beta + rnorm(n, sd = sigma))
data.frame(y_boot = y, x)
}
bootstrap_oracle_se <- function(x, beta, formula, reps) {
sigma <- 1 + 2 * abs(x[, 1])
estimates <- replicate(reps, {
y <- drop(x %*% beta + rnorm(nrow(x), sd = sigma))
coef(lm(formula, data = data.frame(y_boot = y, x)))
})
apply(t(estimates), 2, sd)
}
extract_bootstrap_se <- function(method, fit) {
if (identical(method, "stats::lm")) {
return(as.numeric(fit))
}
if (identical(method, "boot::boot")) {
return(apply(fit$t, 2, sd))
}
if (grepl("^lmboot::", method)) {
return(apply(fit$bootEstParam, 2, sd))
}
as.numeric(fit$std.error)
}
mlxs_lm
Code
lm_results <- list()
lm_grid <- bench_grid[bench_grid$n <= n_sizes[2] & bench_grid$p <= p_sizes[3], ]
lm_accuracy <- list()
for (i in seq_len(nrow(lm_grid))) {
n <- lm_grid$n[i]
p <- lm_grid$p[i]
subset_data <- full_data[1:n, c("y_cont", paste0("x", 1:p))]
lm_formula <- reformulate(paste0("x", 1:p), response = "y_cont")
beta_target <- beta_true[seq_len(p)]
fitters <- list(
"stats::lm" = function() lm(lm_formula, data = subset_data),
"RmlxStats::mlxs_lm" = function() {
fit <- mlxs_lm(lm_formula, data = subset_data)
Rmlx::mlx_eval(fit$coefficients)
fit
},
"fixest::feols" = function() feols(lm_formula, data = subset_data),
"RcppEigen::fastLm" = function() RcppEigen::fastLm(
lm_formula,
data = subset_data
),
"speedglm::speedlm" = function() speedglm::speedlm(
lm_formula,
data = subset_data
)
)
bm <- mark(
"stats::lm" = fitters[["stats::lm"]](),
"RmlxStats::mlxs_lm" = fitters[["RmlxStats::mlxs_lm"]](),
"fixest::feols" = fitters[["fixest::feols"]](),
"RcppEigen::fastLm" = fitters[["RcppEigen::fastLm"]](),
"speedglm::speedlm" = fitters[["speedglm::speedlm"]](),
iterations = 3,
check = function (r1, r2) {
all.equal(as.vector(coef(r1)), as.vector(coef(r2)),
tolerance = 1e-6)
},
filter_gc = FALSE
)
bm$n <- n
bm$p <- p
bm$model_type <- "lm"
lm_results[[i]] <- bm
fits <- lapply(fitters, function(fit_method) fit_method())
scores <- lapply(names(fits), function(method) {
beta_hat <- as.numeric(coef(fits[[method]]))[-1]
data.frame(
model_type = "lm",
n = n,
p = p,
method = method,
accuracy = relative_rmse(beta_hat, beta_target),
stringsAsFactors = FALSE
)
})
lm_accuracy[[i]] <- do.call(rbind, scores)
}
lm_df <- do.call(rbind, lm_results)
all_results <- rbind(all_results, lm_df)
all_accuracy <- rbind(all_accuracy, do.call(rbind, lm_accuracy))
mlxs_glm
Code
glm_results <- list()
glm_grid <- bench_grid[bench_grid$n <= n_sizes[2] & bench_grid$p <= p_sizes[3], ]
glm_accuracy <- list()
for (i in seq_len(nrow(glm_grid))) {
n <- glm_grid$n[i]
p <- glm_grid$p[i]
subset_data <- full_data[1:n, c("y_bin", paste0("x", 1:p))]
glm_formula <- reformulate(paste0("x", 1:p), response = "y_bin")
beta_target <- beta_true[seq_len(p)] / 5
fitters <- list(
"stats::glm" = function() glm(
glm_formula,
family = binomial(),
data = subset_data,
control = list(maxit = 50)
),
"RmlxStats::mlxs_glm" = function() {
fit <- mlxs_glm(
glm_formula,
family = mlxs_binomial(),
data = subset_data,
control = list(maxit = 50, epsilon = 1e-5)
)
Rmlx::mlx_eval(fit$coefficients)
fit
},
"speedglm::speedglm" = function() speedglm::speedglm(
glm_formula,
family = binomial(),
data = subset_data
)
)
bm <- mark(
"stats::glm" = fitters[["stats::glm"]](),
"RmlxStats::mlxs_glm" = fitters[["RmlxStats::mlxs_glm"]](),
"speedglm::speedglm" = fitters[["speedglm::speedglm"]](),
iterations = 3,
check = function (r1, r2) {
all.equal(as.vector(coef(r1)), as.vector(coef(r2)),
tolerance = 1e-4)
},
filter_gc = FALSE
)
bm$n <- n
bm$p <- p
bm$model_type <- "glm"
glm_results[[i]] <- bm
fits <- lapply(fitters, function(fit_method) fit_method())
scores <- lapply(names(fits), function(method) {
beta_hat <- as.numeric(coef(fits[[method]]))[-1]
data.frame(
model_type = "glm",
n = n,
p = p,
method = method,
accuracy = relative_rmse(beta_hat, beta_target),
stringsAsFactors = FALSE
)
})
glm_accuracy[[i]] <- do.call(rbind, scores)
}
glm_df <- do.call(rbind, glm_results)
all_results <- rbind(all_results, glm_df)
all_accuracy <- rbind(all_accuracy, do.call(rbind, glm_accuracy))
mlxs_cv_glmnet
Code
glmnet_results <- list()
glmnet_grid <- bench_grid
glmnet_accuracy <- list()
glmnet_nfolds <- 3L
for (i in seq_len(nrow(glmnet_grid))) {
n <- glmnet_grid$n[i]
p <- glmnet_grid$p[i]
xvars <- paste0("x", 1:p)
x <- X_glmnet[1:n, xvars, drop = FALSE]
y_sparse_subset <- y_sparse[1:n]
beta_target <- beta_glmnet_true[seq_len(p)]
stats_time <- system.time({
fit_stats <- glmnet::cv.glmnet(
x,
y_sparse_subset,
nfolds = glmnet_nfolds
)
as.numeric(fit_stats$lambda.min)
})[["elapsed"]]
mlx_time <- system.time({
fit_mlx <- mlxs_cv_glmnet(
x,
y_sparse_subset,
nfolds = glmnet_nfolds
)
Rmlx::mlx_eval(fit_mlx$glmnet.fit$x_center)
Rmlx::mlx_eval(fit_mlx$glmnet.fit$x_scale)
as.numeric(fit_mlx$lambda.min)
})[["elapsed"]]
bm <- data.frame(
expression = c("glmnet::cv.glmnet", "RmlxStats::mlxs_cv_glmnet"),
median = bench::as_bench_time(c(stats_time, mlx_time)),
stringsAsFactors = FALSE
)
bm$n <- n
bm$p <- p
bm$model_type <- "glmnet"
for (name in setdiff(names(all_results), names(bm))) {
bm[[name]] <- NA
}
bm <- bm[, names(all_results), drop = FALSE]
glmnet_results[[i]] <- bm
fits <- list(
"glmnet::cv.glmnet" = fit_stats,
"RmlxStats::mlxs_cv_glmnet" = fit_mlx
)
scores <- lapply(names(fits), function(method) {
beta_hat <- as.numeric(coef(fits[[method]], s = "lambda.min"))[-1]
data.frame(
model_type = "glmnet",
n = n,
p = p,
method = method,
accuracy = safe_relative_rmse(beta_hat, beta_target),
stringsAsFactors = FALSE
)
})
glmnet_accuracy[[i]] <- do.call(rbind, scores)
}
glmnet_df <- do.call(rbind, glmnet_results)
all_results <- rbind(all_results, glmnet_df)
all_accuracy <- rbind(all_accuracy, do.call(rbind, glmnet_accuracy))
mlxs_prcomp
Code
pca_results <- list()
pca_grid <- bench_grid[bench_grid$n <= n_sizes[2] & bench_grid$p <= p_sizes[4], ]
pca_accuracy <- list()
for (i in seq_len(nrow(pca_grid))) {
n <- pca_grid$n[i]
p <- pca_grid$p[i]
rank_k <- min(8L, max(4L, floor(min(n, p) / 2)))
pca_fixture <- make_pca_fixture(n, p, rank_k)
x <- pca_fixture$x
x_big <- bigstatsr::FBM(nrow = n, ncol = p, init = as.matrix(x))
rotation_true <- pca_fixture$rotation
fitters <- list(
"stats::prcomp" = function() prcomp(
x,
center = TRUE,
scale. = FALSE,
rank. = rank_k
),
"irlba::prcomp_irlba" = function() irlba::prcomp_irlba(
x,
n = rank_k,
center = TRUE,
scale. = FALSE
),
"rsvd::rpca" = function() rsvd::rpca(
x,
k = rank_k,
center = TRUE,
scale = FALSE
),
"bigstatsr::big_randomSVD" = function() bigstatsr::big_randomSVD(
x_big,
fun.scaling = bigstatsr::big_scale(center = TRUE, scale = FALSE),
k = rank_k
),
"RmlxStats::mlxs_prcomp" = function() {
fit <- mlxs_prcomp(
x,
center = TRUE,
scale. = FALSE,
rank. = rank_k,
n_iter = 2,
seed = 1
)
Rmlx::mlx_eval(fit$rotation)
if (!is.null(fit$x)) {
Rmlx::mlx_eval(fit$x)
}
fit
}
)
bm <- mark(
"stats::prcomp" = fitters[["stats::prcomp"]](),
"irlba::prcomp_irlba" = fitters[["irlba::prcomp_irlba"]](),
"rsvd::rpca" = fitters[["rsvd::rpca"]](),
"bigstatsr::big_randomSVD" = fitters[["bigstatsr::big_randomSVD"]](),
"RmlxStats::mlxs_prcomp" = fitters[["RmlxStats::mlxs_prcomp"]](),
iterations = 3,
check = FALSE,
filter_gc = FALSE
)
bm$n <- n
bm$p <- p
bm$model_type <- "pca"
pca_results[[i]] <- bm
fits <- lapply(fitters, function(fit_method) fit_method())
scores <- lapply(names(fits), function(method) {
data.frame(
model_type = "pca",
n = n,
p = p,
method = method,
accuracy = pca_accuracy_score(
fits[[method]],
rotation_true,
pca_fixture$sdev
),
stringsAsFactors = FALSE
)
})
pca_accuracy[[i]] <- do.call(rbind, scores)
}
pca_df <- do.call(rbind, pca_results)
all_results <- rbind(all_results, pca_df)
all_accuracy <- rbind(all_accuracy, do.call(rbind, pca_accuracy))Bootstrap
For bootstrap, we use only smaller datasets due to computational cost.
Code
boot_results <- list()
boot_grid <- bench_grid[bench_grid$n <= n_sizes[2] & bench_grid$p <= p_sizes[2], ]
boot_accuracy <- list()
bootstrap_reps <- if (params$develop) 50L else 100L
oracle_reps <- if (params$develop) 50L else 200L
for (i in seq_len(nrow(boot_grid))) {
n <- boot_grid$n[i]
p <- boot_grid$p[i]
subset_data <- make_bootstrap_data(n, p)
x <- as.matrix(subset_data[paste0("x", seq_len(p))])
beta_target <- beta_true[seq_len(p)]
formula_str <- paste("y_boot ~", paste(paste0("x", 1:p), collapse = " + "))
boot_formula <- as.formula(formula_str)
fit_mlxs <- mlxs_lm(boot_formula, data = subset_data)
oracle_se <- bootstrap_oracle_se(x, beta_target, boot_formula, oracle_reps)
oracle_se <- oracle_se[-1]
boot_stat <- function(dat, idx) {
coef(lm(boot_formula, data = dat[idx, , drop = FALSE]))
}
fitters <- list(
"boot::boot" = function() boot::boot(
subset_data,
statistic = boot_stat,
R = bootstrap_reps,
parallel = "no"
),
"lmboot::paired.boot" = function() lmboot::paired.boot(
boot_formula,
data = subset_data,
B = bootstrap_reps
),
"lmboot::residual.boot" = function() lmboot::residual.boot(
boot_formula,
data = subset_data,
B = bootstrap_reps
),
"mlxs_case" = function() {
summary(
fit_mlxs,
bootstrap = TRUE,
bootstrap_args = list(
B = bootstrap_reps,
seed = 42,
bootstrap_type = "case",
progress = FALSE
)
)
},
"mlxs_resid" = function() {
summary(
fit_mlxs,
bootstrap = TRUE,
bootstrap_args = list(
B = bootstrap_reps,
seed = 42,
bootstrap_type = "resid",
progress = FALSE
)
)
}
)
bm <- mark(
"boot::boot" = fitters[["boot::boot"]](),
"lmboot::paired.boot" = fitters[["lmboot::paired.boot"]](),
"lmboot::residual.boot" = fitters[["lmboot::residual.boot"]](),
"mlxs_case" = {
fit <- fitters[["mlxs_case"]]()
Rmlx::mlx_eval(fit$std.error)
fit
},
"mlxs_resid" = {
fit <- fitters[["mlxs_resid"]]()
Rmlx::mlx_eval(fit$std.error)
fit
},
iterations = 3,
check = FALSE,
filter_gc = FALSE,
memory = FALSE
)
bm$n <- n
bm$p <- p
bm$model_type <- "Bootstrap"
boot_results[[i]] <- bm
fits <- lapply(fitters, function(fit_method) fit_method())
scores <- lapply(names(fits), function(method) {
se_hat <- extract_bootstrap_se(method, fits[[method]])[-1]
data.frame(
model_type = "Bootstrap",
n = n,
p = p,
method = method,
accuracy = relative_rmse(se_hat, oracle_se),
stringsAsFactors = FALSE
)
})
boot_accuracy[[i]] <- do.call(rbind, scores)
}
boot_df <- do.call(rbind, boot_results)
all_results <- rbind(all_results, boot_df)
all_accuracy <- rbind(all_accuracy, do.call(rbind, boot_accuracy))Results: Speed
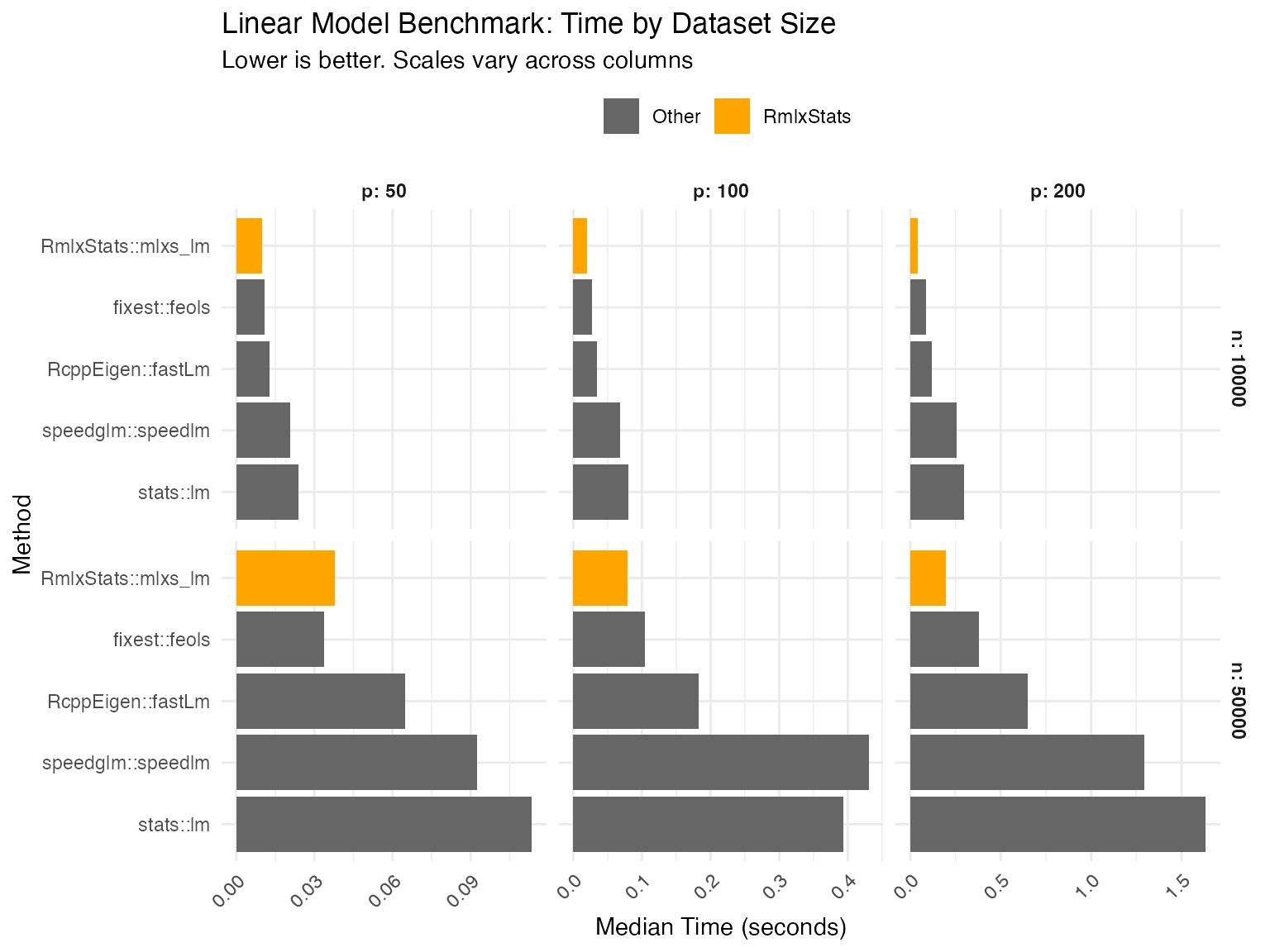
We compare functions both against base R implementation, and against the fastest alternative tested.
Display code
mlxs_lm
Data table
| Method | Median seconds | Rel. to base | Rel. to best |
|---|---|---|---|
| n = 10000, p = 50 | |||
| fixest::feols | 0.010 | 41.5% | 100.0% |
| RcppEigen::fastLm | 0.015 | 59.9% | 144.4% |
| speedglm::speedlm | 0.021 | 85.1% | 205.2% |
| stats::lm | 0.025 | 100.0% | 241.0% |
| RmlxStats::mlxs_lm | 0.012 | 46.2% | 111.4% |
| n = 50000, p = 50 | |||
| fixest::feols | 0.035 | 29.7% | 100.0% |
| RcppEigen::fastLm | 0.088 | 73.9% | 249.1% |
| speedglm::speedlm | 0.11 | 90.6% | 305.1% |
| stats::lm | 0.12 | 100.0% | 336.9% |
| RmlxStats::mlxs_lm | 0.042 | 34.9% | 117.6% |
| n = 10000, p = 100 | |||
| fixest::feols | 0.028 | 31.4% | 128.1% |
| RcppEigen::fastLm | 0.039 | 44.8% | 182.8% |
| speedglm::speedlm | 0.076 | 85.8% | 350.5% |
| stats::lm | 0.088 | 100.0% | 408.4% |
| RmlxStats::mlxs_lm | 0.022 | 24.5% | 100.0% |
| n = 50000, p = 100 | |||
| fixest::feols | 0.10 | 25.1% | 100.0% |
| RcppEigen::fastLm | 0.21 | 51.1% | 203.6% |
| speedglm::speedlm | 0.35 | 84.2% | 335.8% |
| stats::lm | 0.41 | 100.0% | 398.8% |
| RmlxStats::mlxs_lm | 0.12 | 30.2% | 120.4% |
| n = 10000, p = 200 | |||
| fixest::feols | 0.086 | 28.2% | 213.2% |
| RcppEigen::fastLm | 0.12 | 38.7% | 293.3% |
| speedglm::speedlm | 0.26 | 86.0% | 651.0% |
| stats::lm | 0.31 | 100.0% | 756.8% |
| RmlxStats::mlxs_lm | 0.041 | 13.2% | 100.0% |
| n = 50000, p = 200 | |||
| fixest::feols | 0.40 | 26.3% | 196.2% |
| RcppEigen::fastLm | 0.69 | 45.3% | 337.9% |
| speedglm::speedlm | 1.3 | 84.4% | 629.6% |
| stats::lm | 1.5 | 100.0% | 745.9% |
| RmlxStats::mlxs_lm | 0.21 | 13.4% | 100.0% |
| Base is base R implementation in 'stats' or 'boot' packages. | |||
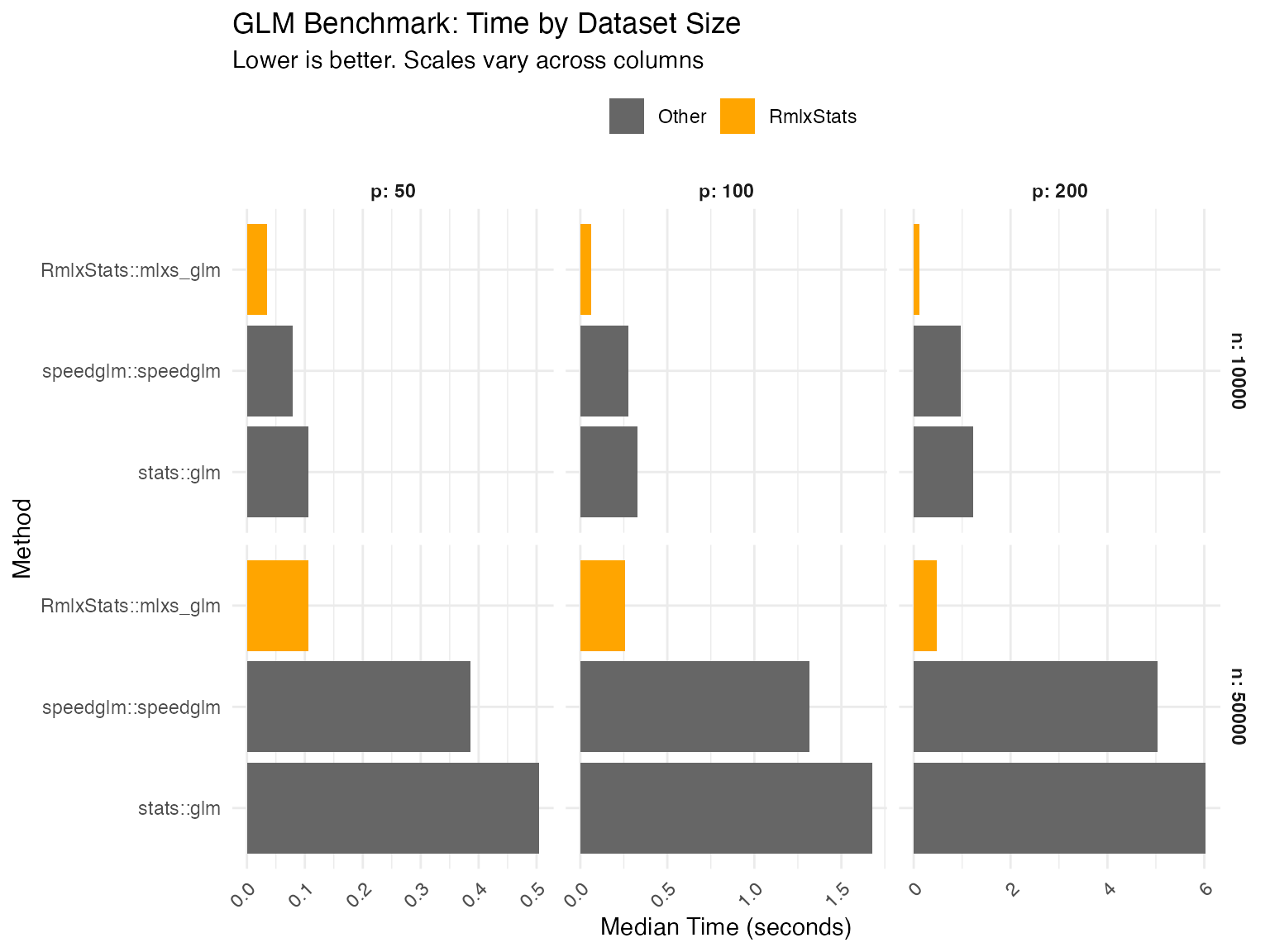
mlxs_glm
Data table
| Method | Median seconds | Rel. to base | Rel. to best |
|---|---|---|---|
| n = 10000, p = 50 | |||
| speedglm::speedglm | 0.080 | 78.0% | 152.5% |
| stats::glm | 0.10 | 100.0% | 195.6% |
| RmlxStats::mlxs_glm | 0.052 | 51.1% | 100.0% |
| n = 50000, p = 50 | |||
| speedglm::speedglm | 0.42 | 81.0% | 254.7% |
| stats::glm | 0.52 | 100.0% | 314.4% |
| RmlxStats::mlxs_glm | 0.16 | 31.8% | 100.0% |
| n = 10000, p = 100 | |||
| speedglm::speedglm | 0.27 | 81.1% | 369.5% |
| stats::glm | 0.34 | 100.0% | 455.7% |
| RmlxStats::mlxs_glm | 0.074 | 21.9% | 100.0% |
| n = 50000, p = 100 | |||
| speedglm::speedglm | 1.4 | 77.0% | 436.7% |
| stats::glm | 1.8 | 100.0% | 567.4% |
| RmlxStats::mlxs_glm | 0.31 | 17.6% | 100.0% |
| n = 10000, p = 200 | |||
| speedglm::speedglm | 1.0 | 82.5% | 726.5% |
| stats::glm | 1.2 | 100.0% | 880.4% |
| RmlxStats::mlxs_glm | 0.14 | 11.4% | 100.0% |
| n = 50000, p = 200 | |||
| speedglm::speedglm | 5.1 | 81.9% | 791.9% |
| stats::glm | 6.2 | 100.0% | 966.4% |
| RmlxStats::mlxs_glm | 0.64 | 10.3% | 100.0% |
| Base is base R implementation in 'stats' or 'boot' packages. | |||
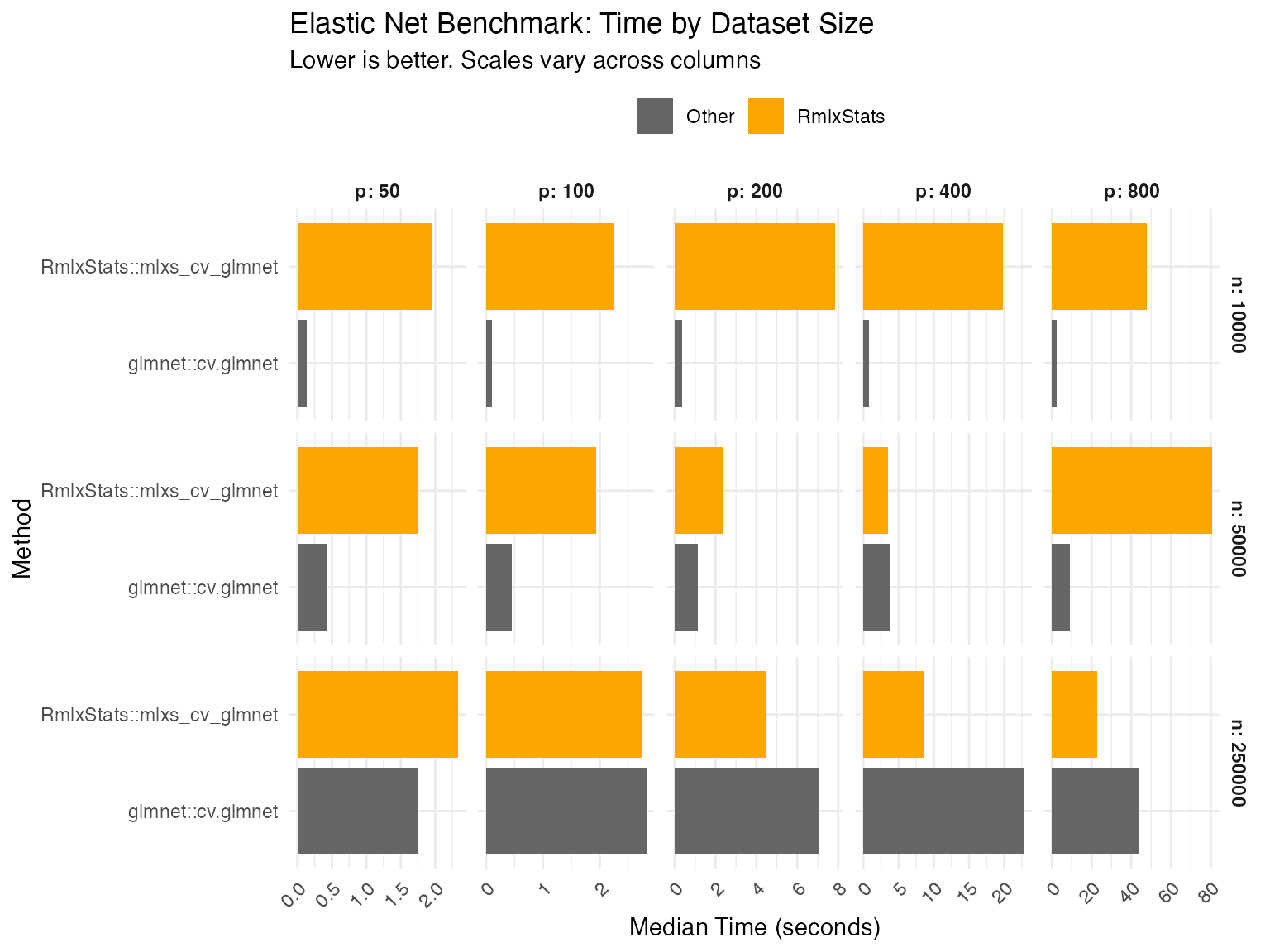
mlxs_cv_glmnet
Data table
| Method | Median seconds | Rel. to base | Rel. to best |
|---|---|---|---|
| n = 10000, p = 50 | |||
| glmnet::cv.glmnet | 0.14 | 100.0% | |
| RmlxStats::mlxs_cv_glmnet | 1.9 | 1348.6% | |
| n = 50000, p = 50 | |||
| glmnet::cv.glmnet | 0.25 | 100.0% | |
| RmlxStats::mlxs_cv_glmnet | 1.7 | 669.7% | |
| n = 250000, p = 50 | |||
| glmnet::cv.glmnet | 1.8 | 100.0% | |
| RmlxStats::mlxs_cv_glmnet | 1.9 | 104.7% | |
| n = 10000, p = 100 | |||
| glmnet::cv.glmnet | 0.16 | 100.0% | |
| RmlxStats::mlxs_cv_glmnet | 1.9 | 1200.0% | |
| n = 50000, p = 100 | |||
| glmnet::cv.glmnet | 0.48 | 100.0% | |
| RmlxStats::mlxs_cv_glmnet | 1.7 | 358.0% | |
| n = 250000, p = 100 | |||
| glmnet::cv.glmnet | 3.3 | 127.1% | |
| RmlxStats::mlxs_cv_glmnet | 2.6 | 100.0% | |
| n = 10000, p = 200 | |||
| glmnet::cv.glmnet | 0.25 | 100.0% | |
| RmlxStats::mlxs_cv_glmnet | 6.0 | 2378.5% | |
| n = 50000, p = 200 | |||
| glmnet::cv.glmnet | 1.4 | 100.0% | |
| RmlxStats::mlxs_cv_glmnet | 2.2 | 157.2% | |
| n = 250000, p = 200 | |||
| glmnet::cv.glmnet | 7.8 | 162.4% | |
| RmlxStats::mlxs_cv_glmnet | 4.8 | 100.0% | |
| n = 10000, p = 400 | |||
| glmnet::cv.glmnet | 0.90 | 100.0% | |
| RmlxStats::mlxs_cv_glmnet | 15. | 1680.8% | |
| n = 50000, p = 400 | |||
| glmnet::cv.glmnet | 4.5 | 130.5% | |
| RmlxStats::mlxs_cv_glmnet | 3.5 | 100.0% | |
| n = 250000, p = 400 | |||
| glmnet::cv.glmnet | 34. | 120.3% | |
| RmlxStats::mlxs_cv_glmnet | 28. | 100.0% | |
| n = 10000, p = 800 | |||
| glmnet::cv.glmnet | 3.2 | 100.0% | |
| RmlxStats::mlxs_cv_glmnet | 30. | 938.0% | |
| n = 50000, p = 800 | |||
| glmnet::cv.glmnet | 9.3 | 100.0% | |
| RmlxStats::mlxs_cv_glmnet | 54. | 586.9% | |
| n = 250000, p = 800 | |||
| glmnet::cv.glmnet | 55. | 234.5% | |
| RmlxStats::mlxs_cv_glmnet | 23. | 100.0% | |
| Base is base R implementation in 'stats' or 'boot' packages. | |||
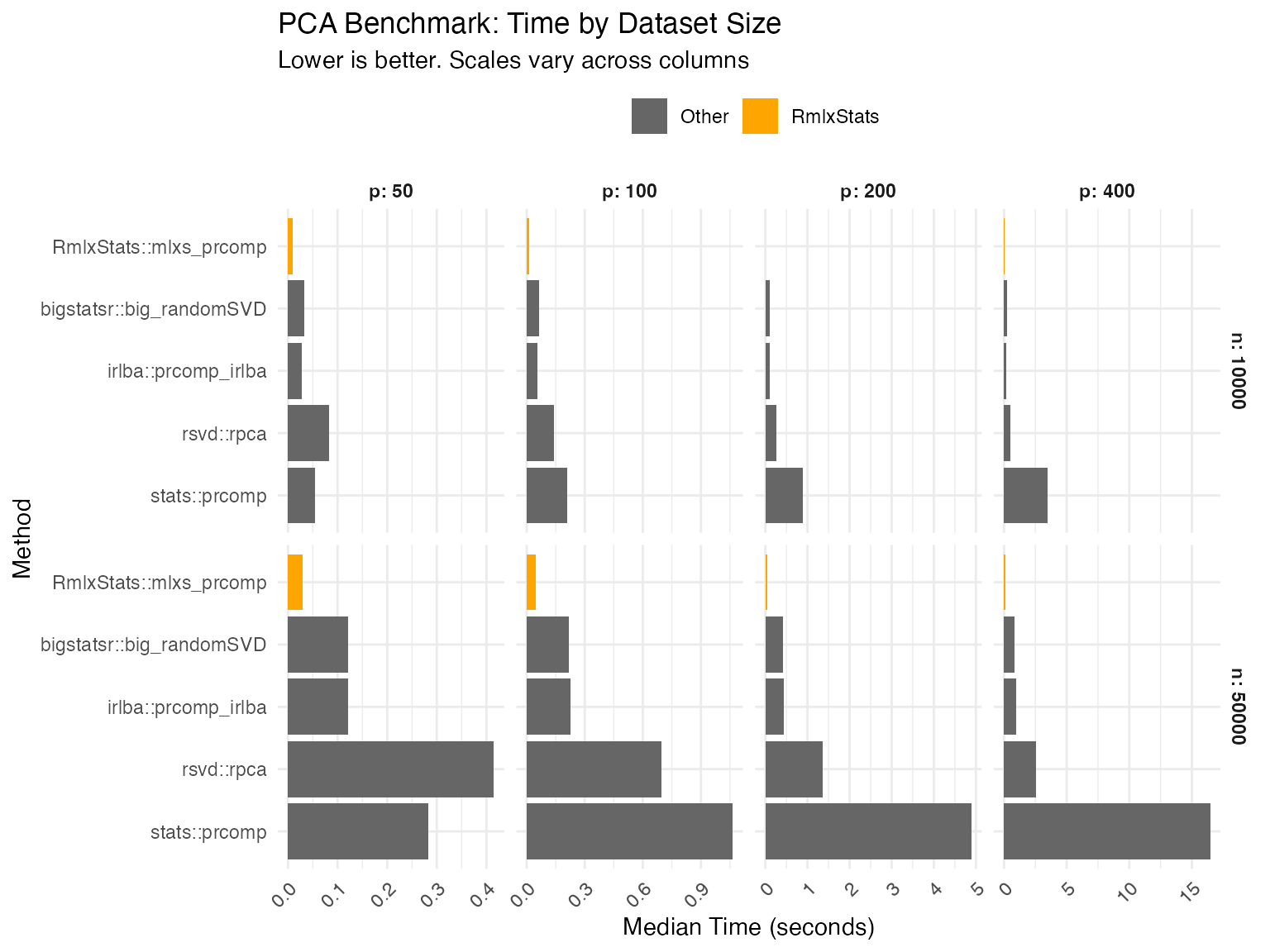
mlxs_prcomp
Data table
| Method | Median seconds | Rel. to base | Rel. to best |
|---|---|---|---|
| n = 10000, p = 50 | |||
| bigstatsr::big_randomSVD | 0.033 | 61.9% | 211.5% |
| irlba::prcomp_irlba | 0.031 | 56.8% | 194.2% |
| rsvd::rpca | 0.084 | 154.7% | 529.0% |
| stats::prcomp | 0.054 | 100.0% | 341.9% |
| RmlxStats::mlxs_prcomp | 0.016 | 29.3% | 100.0% |
| n = 50000, p = 50 | |||
| bigstatsr::big_randomSVD | 0.13 | 41.6% | 588.5% |
| irlba::prcomp_irlba | 0.13 | 42.7% | 604.6% |
| rsvd::rpca | 0.43 | 141.7% | 2006.8% |
| stats::prcomp | 0.30 | 100.0% | 1416.1% |
| RmlxStats::mlxs_prcomp | 0.022 | 7.1% | 100.0% |
| n = 10000, p = 100 | |||
| bigstatsr::big_randomSVD | 0.053 | 23.1% | 397.5% |
| irlba::prcomp_irlba | 0.057 | 25.2% | 433.0% |
| rsvd::rpca | 0.14 | 60.0% | 1029.7% |
| stats::prcomp | 0.23 | 100.0% | 1717.6% |
| RmlxStats::mlxs_prcomp | 0.013 | 5.8% | 100.0% |
| n = 50000, p = 100 | |||
| bigstatsr::big_randomSVD | 0.24 | 21.9% | 849.1% |
| irlba::prcomp_irlba | 0.22 | 20.3% | 785.6% |
| rsvd::rpca | 0.74 | 67.0% | 2594.3% |
| stats::prcomp | 1.1 | 100.0% | 3874.0% |
| RmlxStats::mlxs_prcomp | 0.029 | 2.6% | 100.0% |
| n = 10000, p = 200 | |||
| bigstatsr::big_randomSVD | 0.10 | 11.6% | 645.7% |
| irlba::prcomp_irlba | 0.10 | 11.5% | 641.9% |
| rsvd::rpca | 0.24 | 26.5% | 1473.8% |
| stats::prcomp | 0.90 | 100.0% | 5558.4% |
| RmlxStats::mlxs_prcomp | 0.016 | 1.8% | 100.0% |
| n = 50000, p = 200 | |||
| bigstatsr::big_randomSVD | 0.43 | 10.2% | 484.9% |
| irlba::prcomp_irlba | 0.49 | 11.7% | 557.2% |
| rsvd::rpca | 1.3 | 30.9% | 1465.7% |
| stats::prcomp | 4.2 | 100.0% | 4746.6% |
| RmlxStats::mlxs_prcomp | 0.088 | 2.1% | 100.0% |
| n = 10000, p = 400 | |||
| bigstatsr::big_randomSVD | 0.24 | 6.8% | 640.8% |
| irlba::prcomp_irlba | 0.21 | 6.0% | 566.6% |
| rsvd::rpca | 0.49 | 14.1% | 1342.2% |
| stats::prcomp | 3.5 | 100.0% | 9486.5% |
| RmlxStats::mlxs_prcomp | 0.037 | 1.1% | 100.0% |
| n = 50000, p = 400 | |||
| bigstatsr::big_randomSVD | 0.76 | 4.6% | 793.3% |
| irlba::prcomp_irlba | 0.95 | 5.8% | 990.5% |
| rsvd::rpca | 2.3 | 13.9% | 2368.4% |
| stats::prcomp | 16. | 100.0% | 17068.6% |
| RmlxStats::mlxs_prcomp | 0.096 | 0.6% | 100.0% |
| Base is base R implementation in 'stats' or 'boot' packages. | |||
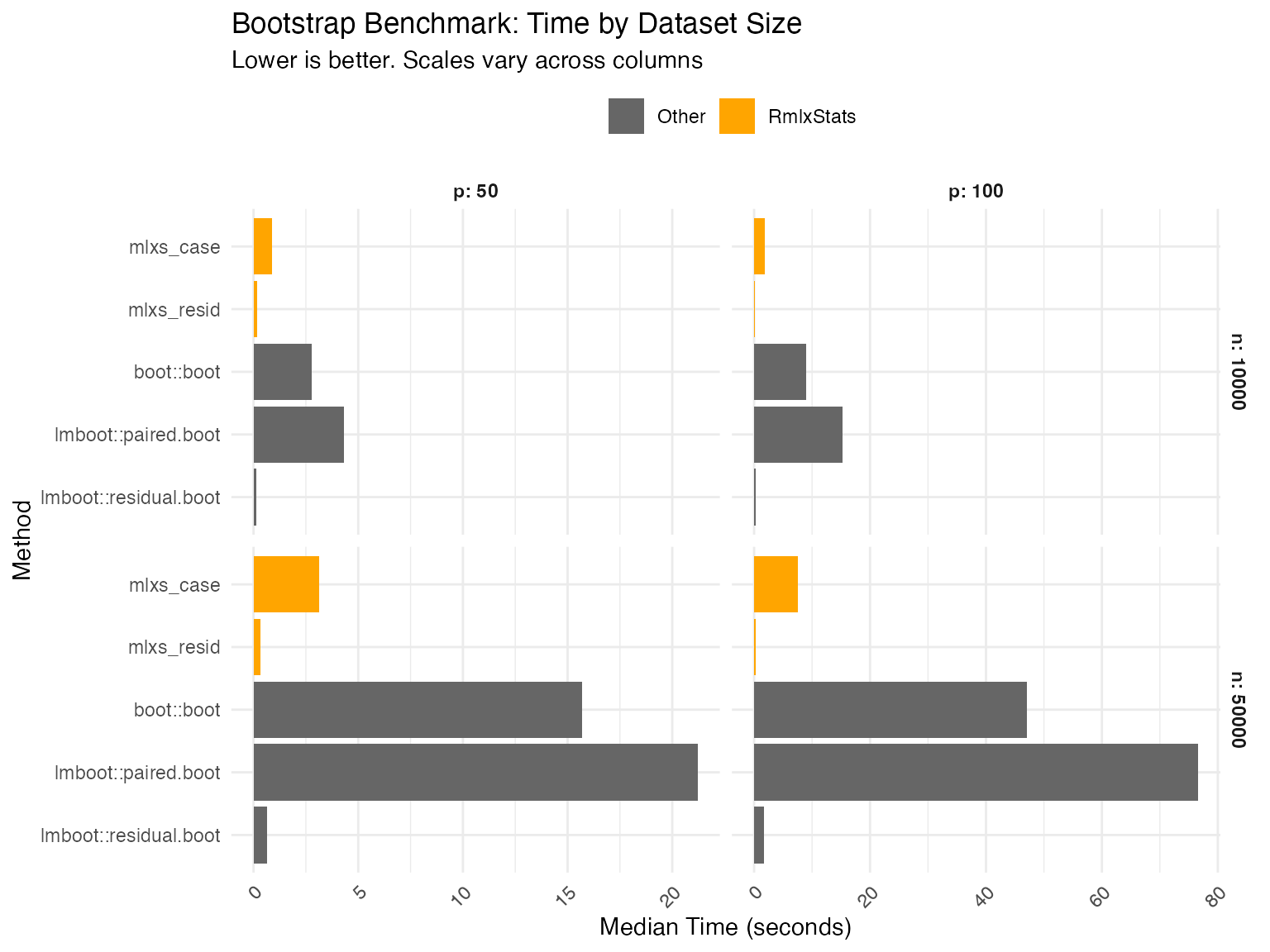
Bootstrap
Data table
| Method | Median seconds | Rel. to base | Rel. to best |
|---|---|---|---|
| n = 10000, p = 50 | |||
| boot::boot | 2.9 | 100.0% | 2094.3% |
| lmboot::paired.boot | 4.4 | 153.9% | 3223.2% |
| lmboot::residual.boot | 0.14 | 4.8% | 100.0% |
| mlxs_case | 1.0 | 36.0% | 754.0% |
| mlxs_resid | 0.22 | 7.7% | 161.9% |
| n = 50000, p = 50 | |||
| boot::boot | 15. | 100.0% | 3523.0% |
| lmboot::paired.boot | 22. | 149.5% | 5267.4% |
| lmboot::residual.boot | 0.63 | 4.2% | 147.9% |
| mlxs_case | 3.2 | 21.6% | 759.8% |
| mlxs_resid | 0.43 | 2.8% | 100.0% |
| n = 10000, p = 100 | |||
| boot::boot | 9.4 | 100.0% | 3670.3% |
| lmboot::paired.boot | 17. | 185.1% | 6793.4% |
| lmboot::residual.boot | 0.38 | 4.0% | 147.1% |
| mlxs_case | 2.1 | 22.0% | 808.8% |
| mlxs_resid | 0.26 | 2.7% | 100.0% |
| n = 50000, p = 100 | |||
| boot::boot | 60. | 100.0% | 10427.6% |
| lmboot::paired.boot | 93. | 154.2% | 16079.0% |
| lmboot::residual.boot | 1.9 | 3.2% | 333.2% |
| mlxs_case | 20. | 33.3% | 3467.2% |
| mlxs_resid | 0.58 | 1.0% | 100.0% |
| Base is base R implementation in 'stats' or 'boot' packages. | |||
Results: Accuracy
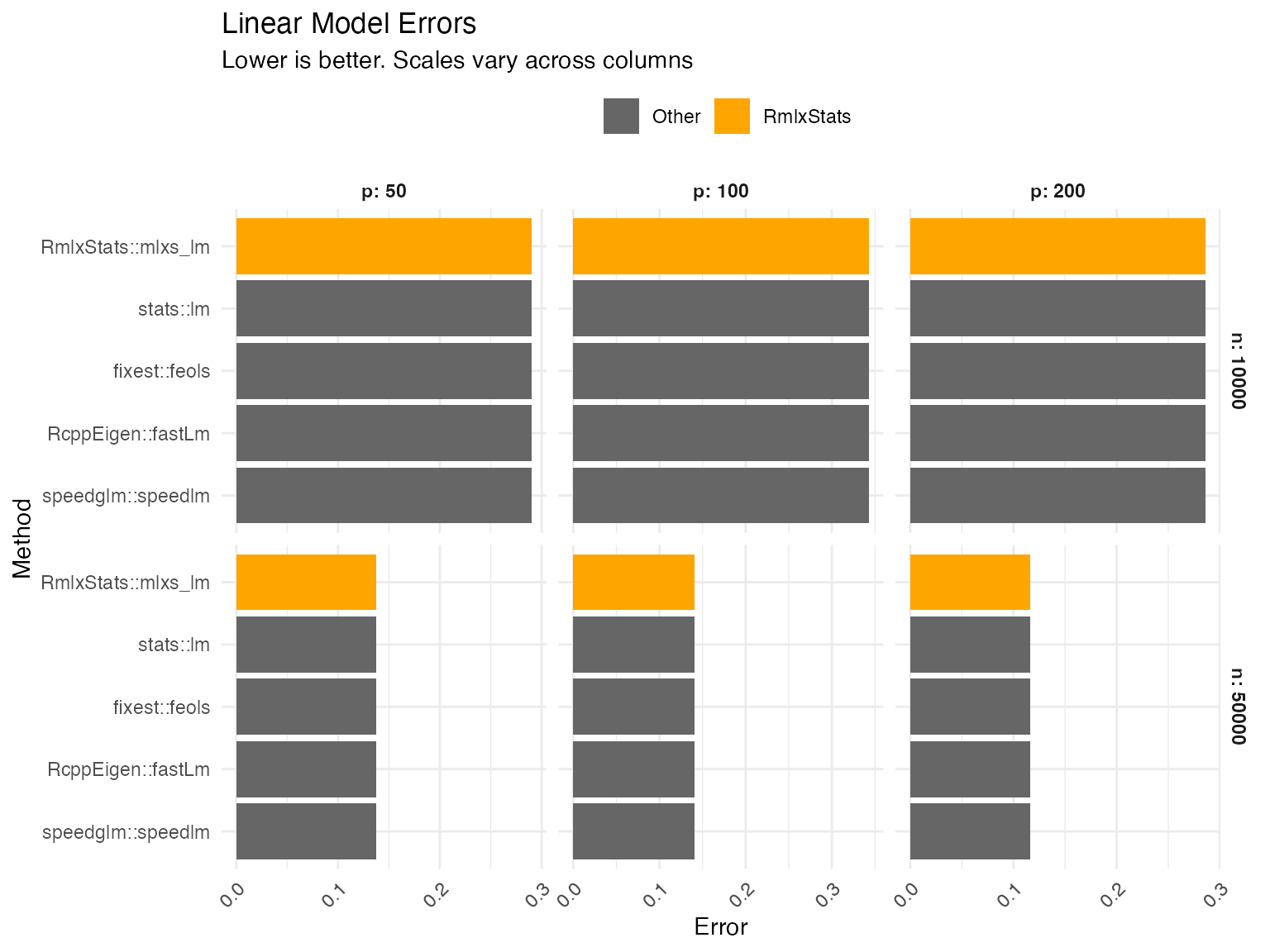
mlxs_lm
Errors are calculated as the relative root mean squared error of the betas (i.e. root mean squared error of the estimated betas compared to the true betas, divided by the root mean square of the true betas).
Data table
| Method | Error | Rel. to best |
|---|---|---|
| n = 10000, p = 50 | ||
| stats::lm | 0.290 | 100.0% |
| RmlxStats::mlxs_lm | 0.290 | 100.0% |
| fixest::feols | 0.290 | 100.0% |
| RcppEigen::fastLm | 0.290 | 100.0% |
| speedglm::speedlm | 0.290 | 100.0% |
| n = 50000, p = 50 | ||
| stats::lm | 0.137 | 100.0% |
| RmlxStats::mlxs_lm | 0.137 | 100.0% |
| fixest::feols | 0.137 | 100.0% |
| RcppEigen::fastLm | 0.137 | 100.0% |
| speedglm::speedlm | 0.137 | 100.0% |
| n = 10000, p = 100 | ||
| stats::lm | 0.343 | 100.0% |
| RmlxStats::mlxs_lm | 0.343 | 100.0% |
| fixest::feols | 0.343 | 100.0% |
| RcppEigen::fastLm | 0.343 | 100.0% |
| speedglm::speedlm | 0.343 | 100.0% |
| n = 50000, p = 100 | ||
| stats::lm | 0.141 | 100.0% |
| RmlxStats::mlxs_lm | 0.141 | 100.0% |
| fixest::feols | 0.141 | 100.0% |
| RcppEigen::fastLm | 0.141 | 100.0% |
| speedglm::speedlm | 0.141 | 100.0% |
| n = 10000, p = 200 | ||
| stats::lm | 0.286 | 100.0% |
| RmlxStats::mlxs_lm | 0.286 | 100.0% |
| fixest::feols | 0.286 | 100.0% |
| RcppEigen::fastLm | 0.286 | 100.0% |
| speedglm::speedlm | 0.286 | 100.0% |
| n = 50000, p = 200 | ||
| stats::lm | 0.116 | 100.0% |
| RmlxStats::mlxs_lm | 0.116 | 100.0% |
| fixest::feols | 0.116 | 100.0% |
| RcppEigen::fastLm | 0.116 | 100.0% |
| speedglm::speedlm | 0.116 | 100.0% |
Graph
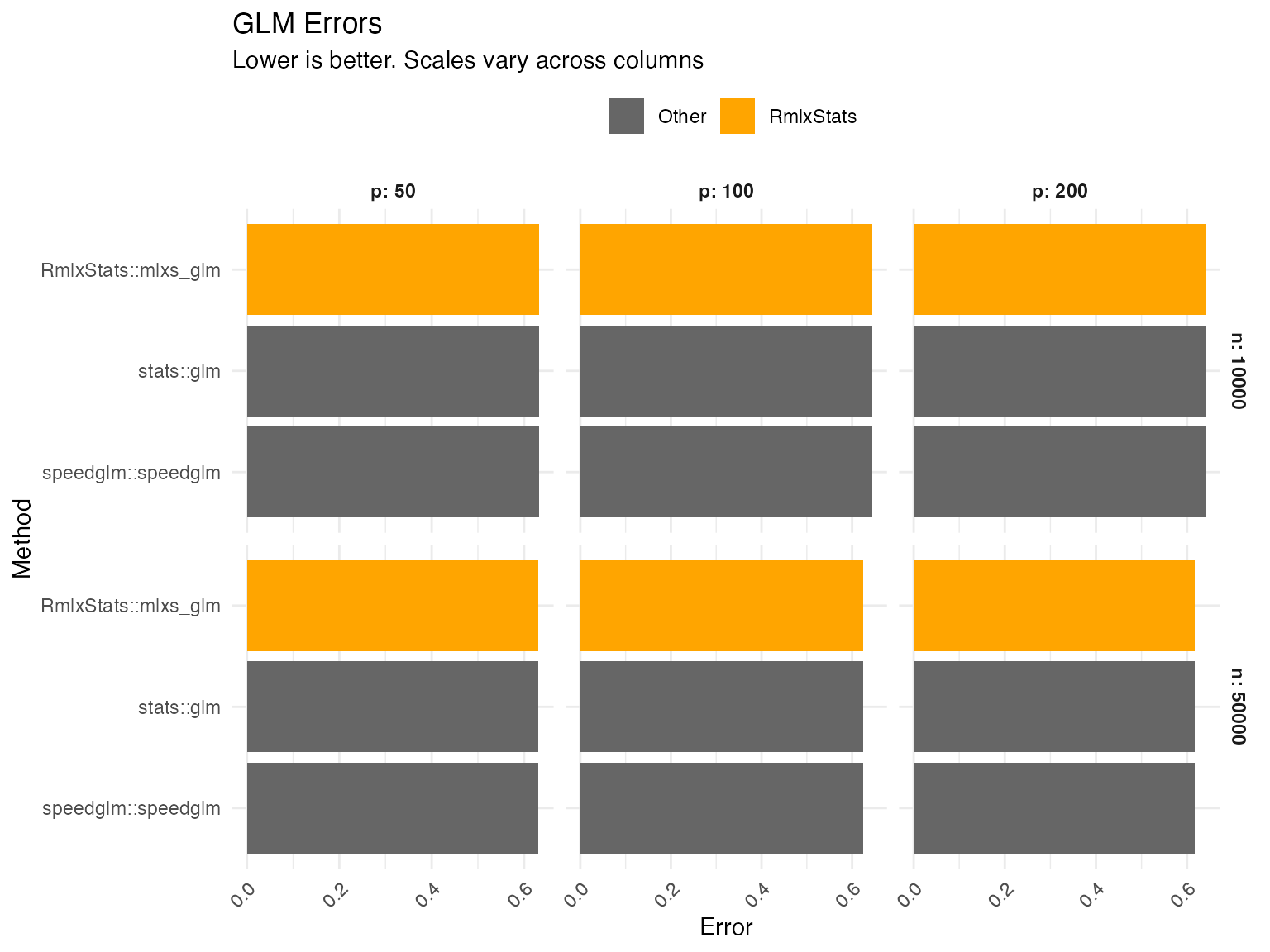
mlxs_glm
Errors are calculated as the relative root mean squared error of the estimated betas.
Data table
| Method | Error | Rel. to best |
|---|---|---|
| n = 10000, p = 50 | ||
| stats::glm | 0.632 | 100.0% |
| RmlxStats::mlxs_glm | 0.632 | 100.0% |
| speedglm::speedglm | 0.632 | 100.0% |
| n = 50000, p = 50 | ||
| stats::glm | 0.630 | 100.0% |
| RmlxStats::mlxs_glm | 0.630 | 100.0% |
| speedglm::speedglm | 0.630 | 100.0% |
| n = 10000, p = 100 | ||
| stats::glm | 0.645 | 100.0% |
| RmlxStats::mlxs_glm | 0.645 | 100.0% |
| speedglm::speedglm | 0.645 | 100.0% |
| n = 50000, p = 100 | ||
| stats::glm | 0.625 | 100.0% |
| RmlxStats::mlxs_glm | 0.625 | 100.0% |
| speedglm::speedglm | 0.625 | 100.0% |
| n = 10000, p = 200 | ||
| stats::glm | 0.641 | 100.0% |
| RmlxStats::mlxs_glm | 0.641 | 100.0% |
| speedglm::speedglm | 0.641 | 100.0% |
| n = 50000, p = 200 | ||
| stats::glm | 0.617 | 100.0% |
| RmlxStats::mlxs_glm | 0.617 | 100.0% |
| speedglm::speedglm | 0.617 | 100.0% |
Graph
mlxs_cv_glmnet
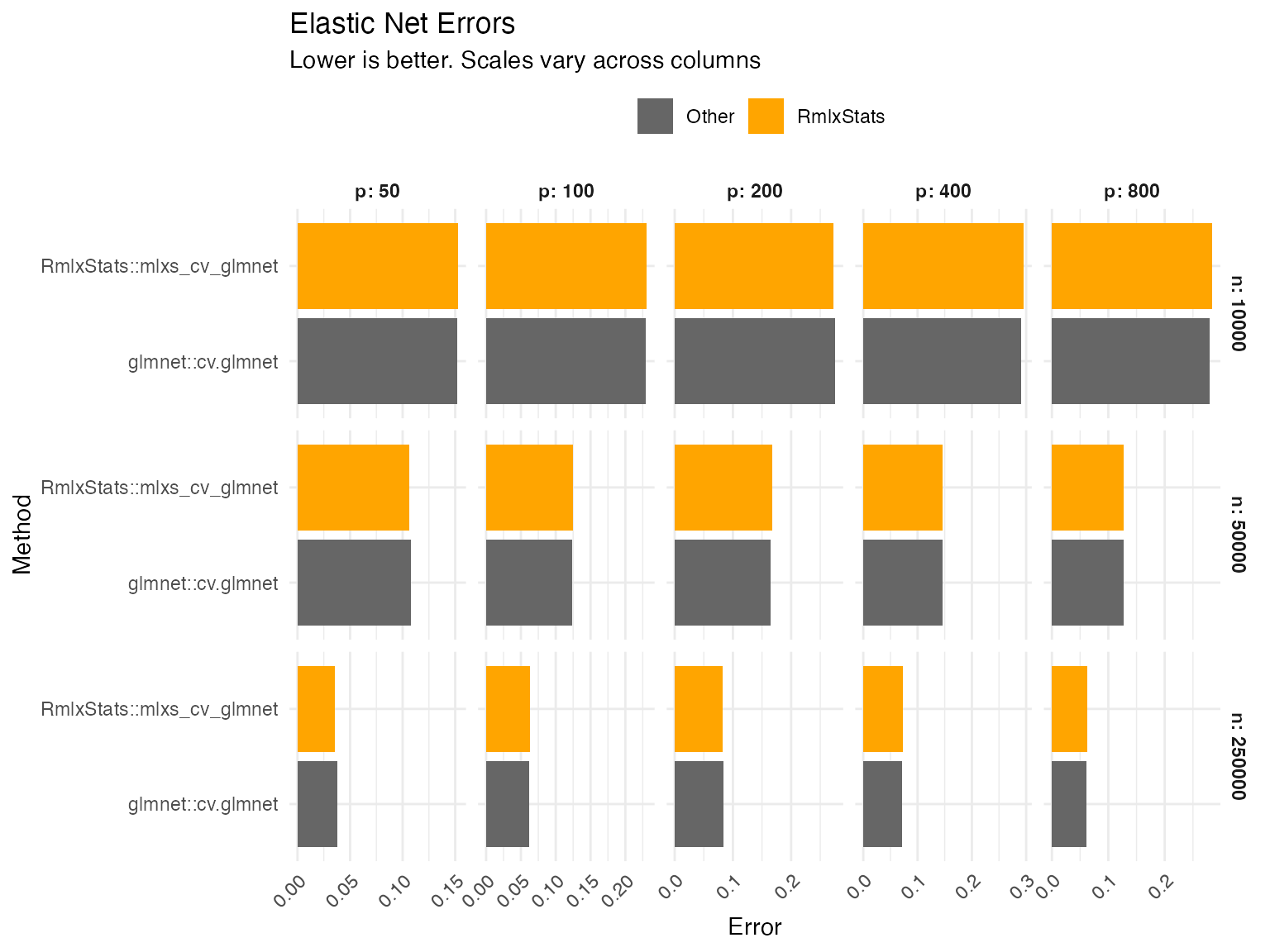
Errors are calculated as the relative root mean squared error of the
estimated betas, for lambda = lambda.min.
Data table
| Method | Error | Rel. to best |
|---|---|---|
| n = 10000, p = 50 | ||
| glmnet::cv.glmnet | 0.152 | 100.0% |
| RmlxStats::mlxs_cv_glmnet | 0.153 | 100.4% |
| n = 50000, p = 50 | ||
| glmnet::cv.glmnet | 0.108 | 101.3% |
| RmlxStats::mlxs_cv_glmnet | 0.106 | 100.0% |
| n = 250000, p = 50 | ||
| glmnet::cv.glmnet | 0.0375 | 105.2% |
| RmlxStats::mlxs_cv_glmnet | 0.0357 | 100.0% |
| n = 10000, p = 100 | ||
| glmnet::cv.glmnet | 0.229 | 100.0% |
| RmlxStats::mlxs_cv_glmnet | 0.230 | 100.6% |
| n = 50000, p = 100 | ||
| glmnet::cv.glmnet | 0.124 | 100.0% |
| RmlxStats::mlxs_cv_glmnet | 0.125 | 100.9% |
| n = 250000, p = 100 | ||
| glmnet::cv.glmnet | 0.0618 | 100.0% |
| RmlxStats::mlxs_cv_glmnet | 0.0634 | 102.6% |
| n = 10000, p = 200 | ||
| glmnet::cv.glmnet | 0.276 | 101.2% |
| RmlxStats::mlxs_cv_glmnet | 0.272 | 100.0% |
| n = 50000, p = 200 | ||
| glmnet::cv.glmnet | 0.164 | 100.0% |
| RmlxStats::mlxs_cv_glmnet | 0.168 | 102.4% |
| n = 250000, p = 200 | ||
| glmnet::cv.glmnet | 0.0832 | 101.1% |
| RmlxStats::mlxs_cv_glmnet | 0.0823 | 100.0% |
| n = 10000, p = 400 | ||
| glmnet::cv.glmnet | 0.291 | 100.0% |
| RmlxStats::mlxs_cv_glmnet | 0.296 | 101.6% |
| n = 50000, p = 400 | ||
| glmnet::cv.glmnet | 0.147 | 100.0% |
| RmlxStats::mlxs_cv_glmnet | 0.147 | 100.0% |
| n = 250000, p = 400 | ||
| glmnet::cv.glmnet | 0.0713 | 100.0% |
| RmlxStats::mlxs_cv_glmnet | 0.0731 | 102.6% |
| n = 10000, p = 800 | ||
| glmnet::cv.glmnet | 0.280 | 100.0% |
| RmlxStats::mlxs_cv_glmnet | 0.284 | 101.8% |
| n = 50000, p = 800 | ||
| glmnet::cv.glmnet | 0.127 | 100.0% |
| RmlxStats::mlxs_cv_glmnet | 0.127 | 100.0% |
| n = 250000, p = 800 | ||
| glmnet::cv.glmnet | 0.0614 | 100.0% |
| RmlxStats::mlxs_cv_glmnet | 0.0622 | 101.3% |
Graph
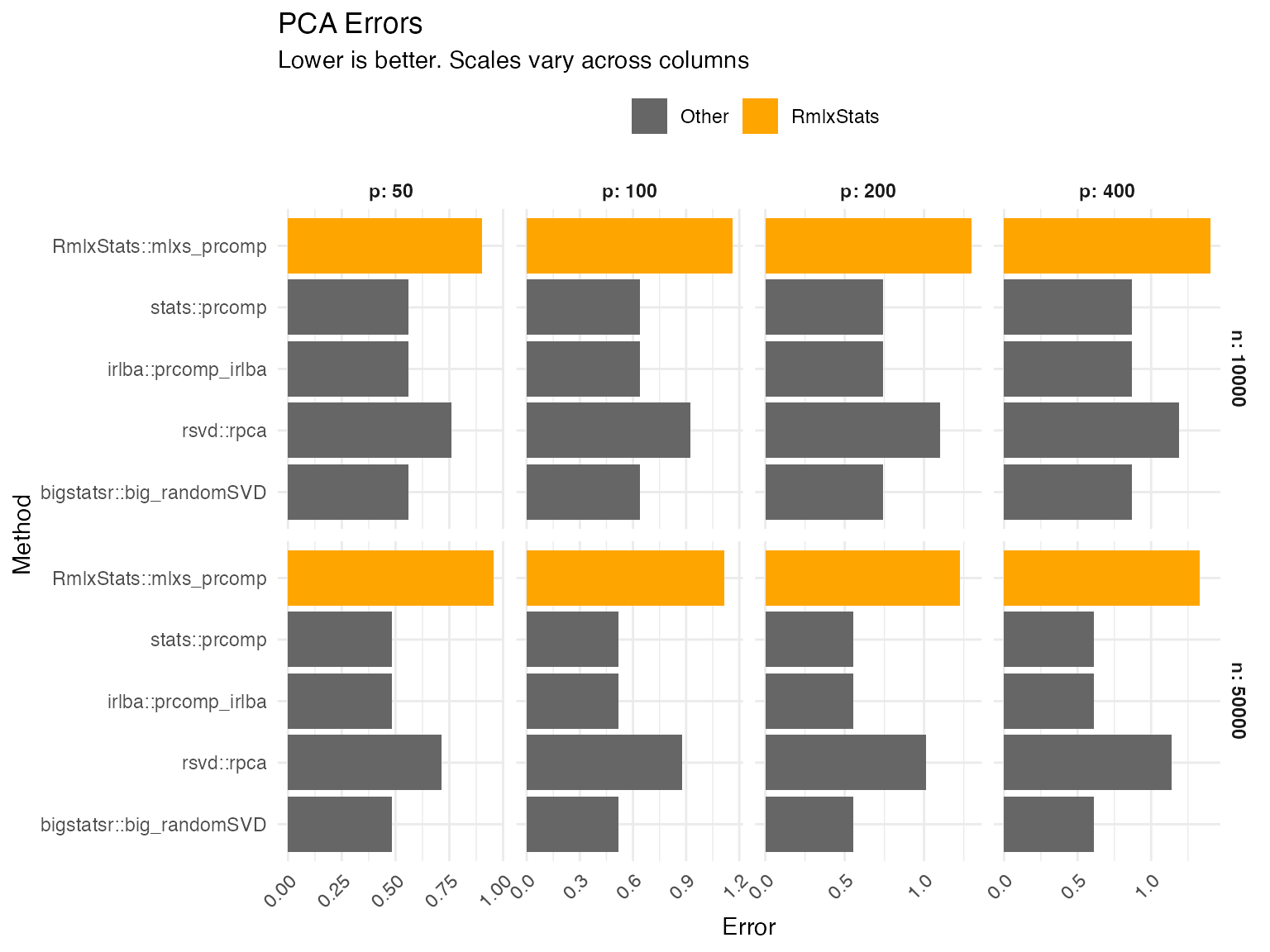
mlxs_prcomp
PCA errors are the projector error of the rotation matrix compared to the true data-generating rotation matrix, plus the relative root mean squared error of the standard deviations. Projector error is equivalent to measuring the principal angles between the estimated and true PCA subspaces.
Data table
| Method | Error | Rel. to best |
|---|---|---|
| n = 10000, p = 50 | ||
| stats::prcomp | 0.559 | 100.0% |
| irlba::prcomp_irlba | 0.559 | 100.0% |
| rsvd::rpca | 0.761 | 136.2% |
| bigstatsr::big_randomSVD | 0.559 | 100.0% |
| RmlxStats::mlxs_prcomp | 0.900 | 161.0% |
| n = 50000, p = 50 | ||
| stats::prcomp | 0.483 | 100.0% |
| irlba::prcomp_irlba | 0.483 | 100.0% |
| rsvd::rpca | 0.712 | 147.5% |
| bigstatsr::big_randomSVD | 0.483 | 100.0% |
| RmlxStats::mlxs_prcomp | 0.957 | 198.4% |
| n = 10000, p = 100 | ||
| stats::prcomp | 0.642 | 100.0% |
| irlba::prcomp_irlba | 0.642 | 100.0% |
| rsvd::rpca | 0.924 | 144.0% |
| bigstatsr::big_randomSVD | 0.642 | 100.0% |
| RmlxStats::mlxs_prcomp | 1.16 | 181.3% |
| n = 50000, p = 100 | ||
| stats::prcomp | 0.516 | 100.0% |
| irlba::prcomp_irlba | 0.516 | 100.0% |
| rsvd::rpca | 0.875 | 169.6% |
| bigstatsr::big_randomSVD | 0.516 | 100.0% |
| RmlxStats::mlxs_prcomp | 1.11 | 215.7% |
| n = 10000, p = 200 | ||
| stats::prcomp | 0.740 | 100.0% |
| irlba::prcomp_irlba | 0.740 | 100.0% |
| rsvd::rpca | 1.10 | 148.8% |
| bigstatsr::big_randomSVD | 0.740 | 100.0% |
| RmlxStats::mlxs_prcomp | 1.30 | 175.5% |
| n = 50000, p = 200 | ||
| stats::prcomp | 0.552 | 100.0% |
| irlba::prcomp_irlba | 0.552 | 100.0% |
| rsvd::rpca | 1.01 | 183.4% |
| bigstatsr::big_randomSVD | 0.552 | 100.0% |
| RmlxStats::mlxs_prcomp | 1.23 | 222.3% |
| n = 10000, p = 400 | ||
| stats::prcomp | 0.870 | 100.0% |
| irlba::prcomp_irlba | 0.870 | 100.0% |
| rsvd::rpca | 1.19 | 136.7% |
| bigstatsr::big_randomSVD | 0.870 | 100.0% |
| RmlxStats::mlxs_prcomp | 1.40 | 161.1% |
| n = 50000, p = 400 | ||
| stats::prcomp | 0.611 | 100.0% |
| irlba::prcomp_irlba | 0.611 | 100.0% |
| rsvd::rpca | 1.14 | 186.0% |
| bigstatsr::big_randomSVD | 0.611 | 100.0% |
| RmlxStats::mlxs_prcomp | 1.33 | 217.6% |
Graph
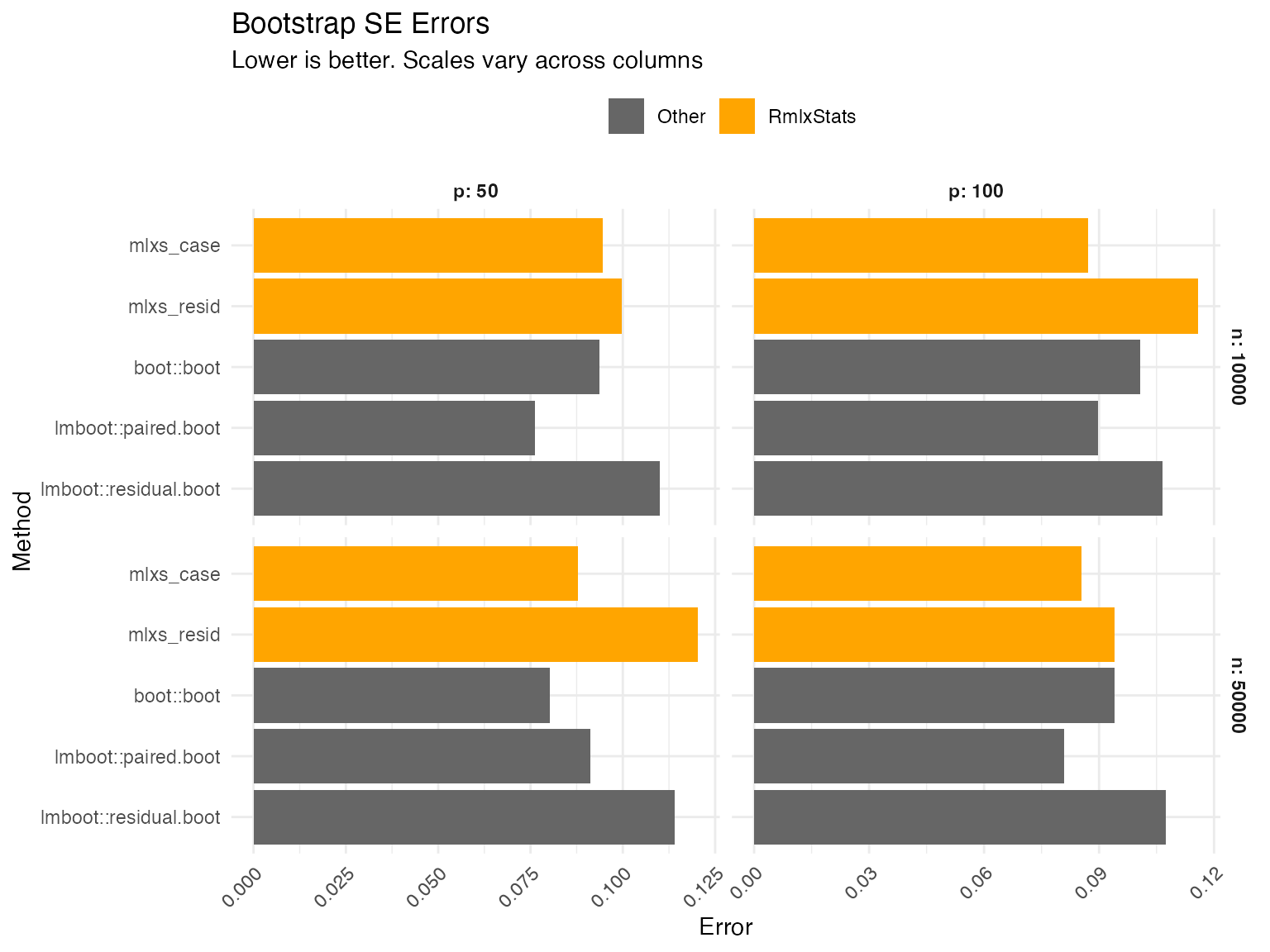
Bootstraps
Errors are calculated as the root mean squared error of the bootstrap-estimated standard errors. We calculate the true standard error by repeatedly estimating linear models, drawing a new dependent variable from the (known) data generating process and calculating the standard deviation of the estimates.
Data table
| Method | Error | Rel. to best |
|---|---|---|
| n = 10000, p = 50 | ||
| boot::boot | 0.0937 | 123.0% |
| lmboot::paired.boot | 0.0761 | 100.0% |
| lmboot::residual.boot | 0.110 | 144.6% |
| mlxs_case | 0.0945 | 124.1% |
| mlxs_resid | 0.0997 | 130.9% |
| n = 50000, p = 50 | ||
| boot::boot | 0.0803 | 100.0% |
| lmboot::paired.boot | 0.0912 | 113.6% |
| lmboot::residual.boot | 0.114 | 141.9% |
| mlxs_case | 0.0879 | 109.4% |
| mlxs_resid | 0.120 | 149.7% |
| n = 10000, p = 100 | ||
| boot::boot | 0.101 | 115.7% |
| lmboot::paired.boot | 0.0897 | 103.0% |
| lmboot::residual.boot | 0.106 | 122.3% |
| mlxs_case | 0.0870 | 100.0% |
| mlxs_resid | 0.116 | 133.1% |
| n = 50000, p = 100 | ||
| boot::boot | 0.0940 | 116.1% |
| lmboot::paired.boot | 0.0810 | 100.0% |
| lmboot::residual.boot | 0.107 | 132.8% |
| mlxs_case | 0.0854 | 105.5% |
| mlxs_resid | 0.0940 | 116.1% |
Graph